DNA in eukaryotic cells is normally found associated with protiens called histone in particles called nucleosomes linked by short segments of DNA. Chromatin remodelers are specialized ATP-dependent molecular machines that can reorganize the structure of nucelsomes as needed for such processes such as replication, transcription and DNA repair.
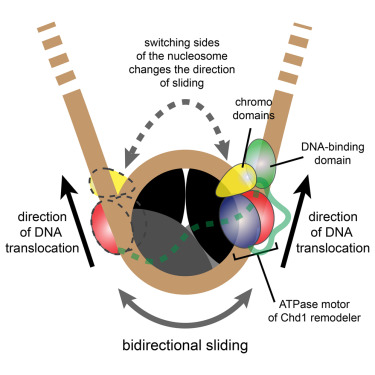
Remodeling results in evenly spaced nucleosomes along the DNA strand yet the way remodelers achieve this is not understood. CHD-1 is a remodeler important for transcription. Here, the authors show that the Chd1 remodeler shifts DNA back and forth by dynamically alternating between different segments of the nucleosome. During sliding, Chd1 generates unstable remodeling intermediates that spontaneously relax to a pre-remodeled position. They demonstrate that nucleosome sliding is tightly controlled by two regulatory domains: the DNA binding domain, which interferes with sliding when its range is limited by a truncated linking segment, and the chromodomains, which play a key role in substrate discrimination. They propose that active interplay of the ATPase motor with the regulatory domains may promote dynamic nucleosome structures uniquely suited for histone exchange and chromatin reorganization during transcription. This work advances our understanding of the Chd1 chromatin remodeler, and puts forward several concepts that may also apply to other remodeling enzymes.
See: Yupeng Qiu, Rob Levendosky, Srinivas Chakravarthy, Ashok Patel, Gregory Bowman, Sua Myong. Molecular Cell 68(1):76-88.
While most of the study was based on single molecule FRET, high quality SEC-SAX at BioCAT played a key role in testing hypothetical configurations of mutant proteins generated to test the proposed packing models.