Science Highlights
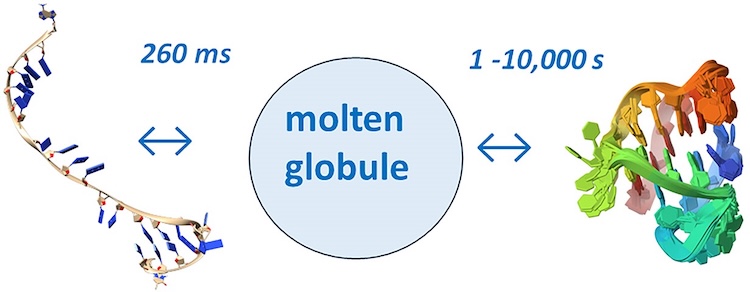
Unraveling folding pathways of dynamic DNA quadruplexes
G-quadruplexes (G4s) are non-B form DNA structures containing genomic regions rich in guanine that sometimes fold into and often act as transcriptional regulators. They are frequently located at regulatory sites including promoters, replication origins and telomeres11. Understanding their folding pathways, while important to a fundamental understanding of form and function, has been challenging. G4 quadruplex folding pathways are distinct from not only proteins but also duplex RNA and DNA12. Previous work characterizing folding pathways using FRET, CD or stopped-flow absorbance suggested a complex, multi-step pathway. Researchers from the University of Louisville, some of whom are frequent BioCAT users, performed continuous-flow mixing time-resolved SAXS experiments to directly structurally characterize, for the first time, early steps in the G4 quadruplex folding pathway.
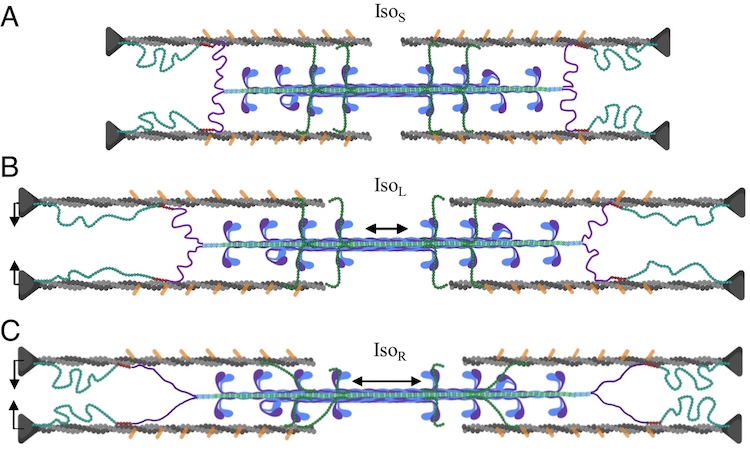
A role for titin in residual force enhancement in skeletal muscle
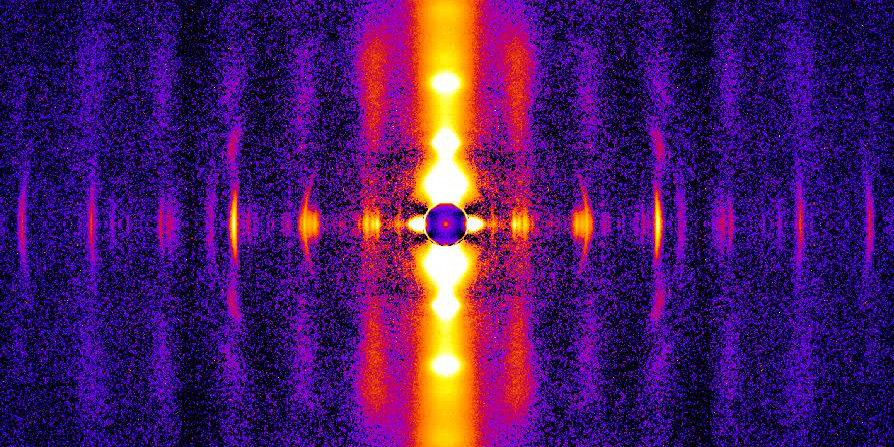
Residual force enhancement (RFE) is a property of skeletal muscle where more force is produced after an active stretch than if it were simply activated at the longer length. This property is important for jumping, locomotive, and stabilizing movements of the body. The molecular mechanism underlying RFE is not well understood. In muscle, the protein titin connects myofilaments and has been shown to have many roles in sarcomere stability and modulating contraction. The authors of this study investigated titin’s function during contraction using small-angle X-ray diffraction of untreated WT mouse skeletal muscle and muscle where 50% of the titin was cleaved. They compared their results to those obtained from mdm titin mutant mce that do not show residual force enhancement.
Role of titin in the pathophysiology of diaphragm weakness during mechanical ventilation
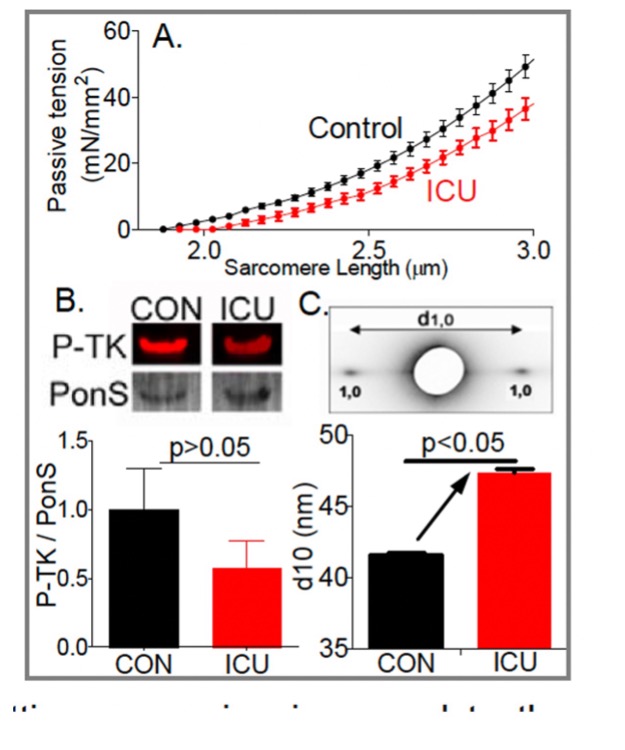
Diaphragm unloading during mechanical ventilation is an important clinical problem. The diaphragm is the main muscle of respiration and contracts during each breath, thereby changing the anatomic configurations of the chest wall so that air flows into the lungs. Increased diaphragm loading is associated with diaphragm fiber contractile dysfunction, atrophy and injury. Whereas these effects of increased loading on the diaphragm take months or years to develop, the effects of decreased loading, as occurs when ICU patients are mechanically ventilated, occur extremely rapidly, within hours. Not known is which structures sense the mechanical unloading of the diaphragm and set in motion the molecular cascades leading to atrophy. Here we test the hypothesis that the mechano-sensor protein is titin, a giant elastic protein connecting Z-disks and the thick filaments in the A-band.
News

Beamline scientist position open at BioCAT
BioCAT is seeking a Ph.D. level scientist to lead the X-ray Fiber Diffraction Program at the beamline with a major emphasis on studies of striated muscle systems.

BioCAT wins NIH grant renewal
We are pleased to announce that the P30 grant that funds BioCAT has been renewed by the NIH and we are looking forward to continuing to serve this community and generate exciting science.
Everything BioSAXS 11 Workshop Wrap-Up
BioCAT held its eleventh intensive HOW-TO course in BioSAXS from 10/7/25-10/10/25 with 35 remote participants. There were four days of lectures and hands-on software tutorials on the basics of BioSAXS data collection and processing from expert practitioners in the field. Participants could elect to mail in samples for data collection prior to the course, and roughly a quarter of them sent research samples and were able to analyze their own data as part of the workshop.