On average, one hundred billion cells in the human body divide over the course of a day. Most of the time the body gets it right but sometimes problems in cell replication can lead to abnormalities in chromosomes—-resulting in many types of disorders from cancer to Down syndrome. Now, researchers from the University of Pennsylvania School of Medicine (UPSM) have defined the structure of a key molecule that plays a central role in how DNA is duplicated and then moved correctly and equally into two daughter cells to produce two exact copies of the mother cell. Without this molecule, entire chromosomes could be lost during cell division, so this work is a major advance in understanding the molecules driving human genetic inheritance. Two U.S. Department of Energy x-ray light sources, including the Advanced Photon Source (APS) at Argonne National Laboratory, were important tools for the researchers carrying out this study.
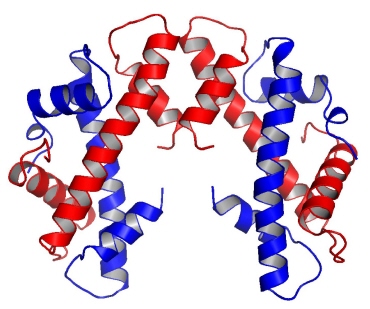
The UPSM researchers report, in the September 16, 2010 issue of Nature, the structure of the CENP-A molecule, which defines a part of the chromosome called the centromere. This is a constricted area to which specialized molecules called spindle fibers attach in order to help pull daughter cells apart during cell division.
“Our work gives us the first high-resolution view of the molecules that control genetic inheritance at cell division,” said Ben Black, assistant professor of Biochemistry and Biophysics at UPSM and a coauthor of the study. “This is a big step forward in a puzzle that biologists have been chipping away at for over 150 years.”
Investigators have known for the last 15 years that part of cell division is controlled by epigenetic processes, the series of actions that affect the protein spools around which DNA is tightly bound, rather than encoded in the DNA sequence itself. Those spools are built of histone proteins, and chemical changes to these spool proteins can either loosen or tighten their interaction with DNA. Epigenetics alter the readout of the genetic code, in some cases ramping a gene’s expression up or down. In the case of the centromere, it marks the site where spindle fibers attach independently of the underlying DNA sequence. CENP-A has been suspected to be the key epigenetic marker protein.
However, what hasn’t been known is how CENP-A epigenetically marks the centromere to direct inheritance. In this study, the UPSM team found the structural features that confer CENP-A the ability to mark centromere location on each chromosome.
This is important because without CENP-A or the centromere mark it creates, the entire chromosome—and all of the genes it houses—are lost at cell division.
Utilizing the Biophysics Collaborative Access Team (BioCAT) 18-ID and General Medicine and Cancer Institutes Collaborative Access Team (GM/CA-CAT) 23-ID-D beamlines at the APS, and the 8.2.2 beamline at the Advanced Light Source at Lawrence Berkeley National Laboratory, the group solved CENP-A’s structure to determine how it specifically marks the centromere on each chromosome and surmise from that how the epigenetic mark is copied correctly in each cell division. They found that CENP-A changes the shape of the nucleosome of which it’s a part, also making it more rigid than other nucleosomes without CENP-A. The nucleosome is the combination of DNA wound around a histone protein core—the DNA thread wrapped around the histone spool. The CENP-A nucleosome is copied several times to create a unique epigenetic area, different from the rest of the chromosome. CENP-A replaces histone H3 in the nucleosomes located at the centromere.
This CENP-A centromere identifier attracts other proteins, and in cell division builds a massive structure, the kinetochore, for pulling the duplicated chromosomes apart during cell division.
Besides being a major advance in the understanding of molecules driving human inheritance, this work also brings about the exciting prospect that the key epigenetic components are now in hand to engineer clinically useful artificial chromosomes that will be inherited alongside our own natural chromosomes— and with the same high fidelity, says Black.
See: Nikolina Sekulic, Emily A. Bassett, Danielle J. Rogers, and Ben E. Black, “The structure of (CENP-A–H4)2 reveals physical features that mark centromeres,” Nature 467, 347 (16 September 2010). DOI:10.1038/nature09323
This work was supported by the National Institutes of Health (NIH) research grant GM82989, a Career Award in the Biomedical Sciences from the Burroughs Wellcome Fund, and a Rita Allen Foundation Scholar Award to B.E.B. N.S. is supported by a postdoctoral fellowship from the American Cancer Society and E.A.B. has been supported by the Penn Structural Biology Training Grant (NIH GM08275) and a predoctoral fellowship from the American Heart Association. Use of the APS was supported by the DOE-BES, under Contract No. DE-AC02-06CH11357.
The original University of Pennsylvania School of Medicine press release is no longer available online.