
You are what you eat” is just another way of saying that input determines output, after some metabolic messing around in between. It’s the “in between” that interests biologists, because that is where the difference between healthy and diseased cells can originate. If the normal sequence of intermediate steps between the beginning and end of a biochemical pathway is disrupted, the disruption can lead to major changes in cellular functioning. A better understanding of the input-output relationships is critical to medical progress. The steps necessary for producing the right output can be numerous and complex, and some pathways have taken decades to unravel. In one of the identified mechanisms, called “two-component signal transduction,” the input causes a receiver domain to be phosphorylated, a step that governs what happens to the output modules. By employing x-ray scattering and electron microscopy researchers from Pennsylvania State University; the University of California, Berkeley; Lawrence Berkeley National Laboratory; The University of Georgia; and the Illinois Institute of Technology using the BioCAT 18-ID beamline at the APS were able to describe—in stunning detail—a novel two-component mechanism for assembling a protein associated with bacterial transcription. Their work greatly advances our understanding of what happens in normal and, by inference, diseased cells.

Transcription is the process by which the cell’s DNA is read to make RNA, which is in turn translated into protein. The DNA-to-RNA transition is, naturally, quite complex and intricately regulated, because the cell can only function normally if DNA is translated to protein at the exact right moment. RNA polymerases—those enzymes responsible for creating RNA from DNA—are many and varied. One RNA polymerase will function normally only with the help of enhancer-binding proteins, such as the nitrogen-regulatory protein C of enteric bacteria (NtrC). NtrC is believed to be involved in regulating transcription for up to 2% of the genome, so determining its structure is of great importance.
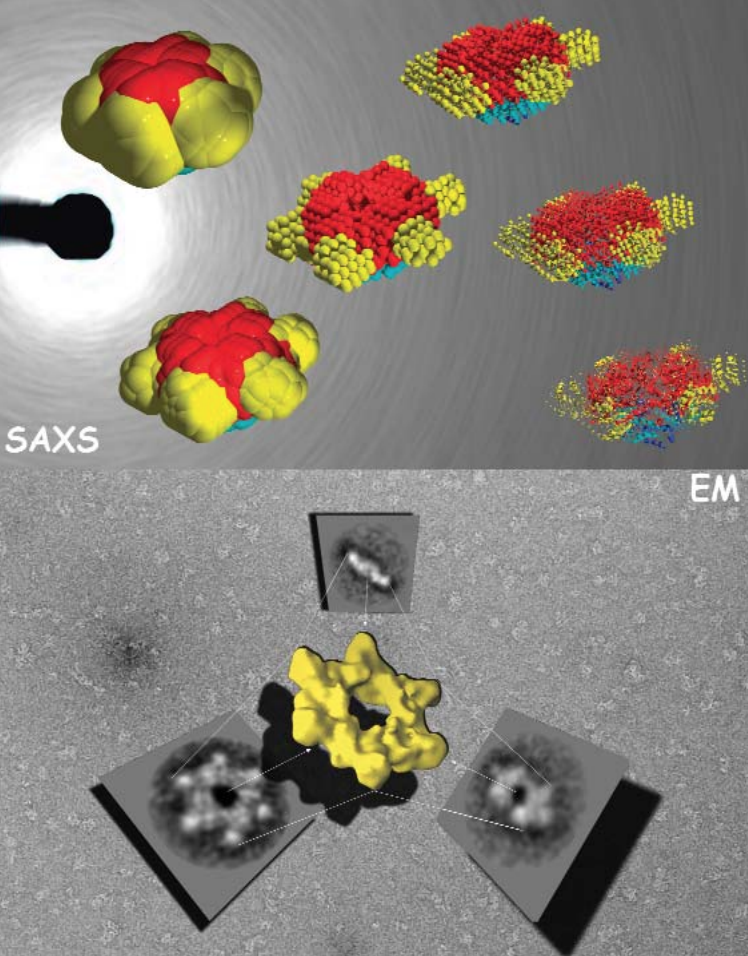
The structures of the enhancer-binding proteins like NtrC are quite complex—in addition to their ATPase and DNA-binding domains, they have a regulatory domain, which (in half the cases) has been found to be a receiver domain for a two-component signal transduction system. Until now, available structural data on NtrC - like proteins, while suggestive of how the “in between” steps could occur, did not provide a complete model. The researchers in this study, with the help of beamline 18-ID, used small- and wide-angle x-ray scattering (SAXS/WAXS) and electron microscopy (EM) to develop structures for the full - length NtrC from Salmonella typhimurium. These new data not only provide a complete picture of how the individual domains of the protein fit into the activated enzyme, but they also reveal a new mechanism for using two-component signal transduction in regulating AAA+ ATPase domains. The group was also able to identify additional structural features — the ordering of the DNA binding domain and an outward extension of a GAFTGA loop region—that could be important in the hydrolysis-activation process and may suggest avenues for further study.
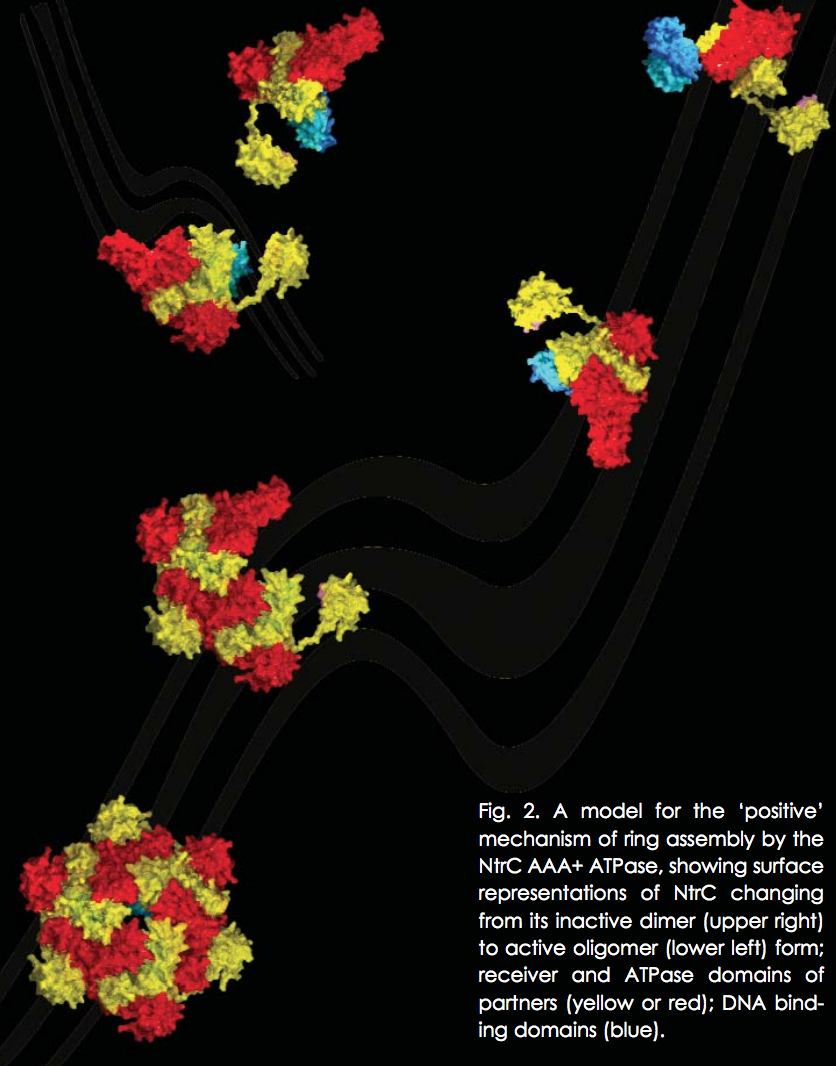
The new mechanism discovered by the research group involves juxtaposition of the receiver domains and the ATPase ring. The beautiful structure of the NtrC revealed by this study shows a central ATPase domain, with six receiver domains packed tightly around it (Fig. 1). Three dimers of DNA-binding domains lie underneath the main ring—these domains become flexible and are believed to detach from the central ring when inorganic phosphate is released. In addition, by using the NtrC structure and previous biochemical data, the researchers were able to postulate contact between the activated receiver domain of one subunit and the ATPase domain of another, thus explaining how the NtrC receiver domains play a positive role in regulating assembly of the ATPase domains into their functional ring form (Fig. 2). This configuration of receiver and ATPase domains differs markedly from previous models proposed for how two-component signal transduction “negatively” regulates assembly of AAA+ ATPase rings in a related protein called NtrC1. By using the new model, researchers can identify structural differences underlying positive versus negative regulation for this family of enhancer-binding proteins. The structure also allows hypotheses about how specific changes in NtrC amino acids and in order-disorder of the GAFTGA loop and DNA-binding domains could affect assembly, thereby suggesting explanations for disease states, drug-design possibilities, and, in the broad view, how input affects output.
See: S. De Carlo, B. Chen, T.R. Hoover, E. Kondrashkina, E. Nogales, and B.T. Nixon, “The Structural Basis for Regulated Assembly and Function of the Transcriptional Activator NtrC,” Gene. Dev. 20, 1485 (2006, cover story). DOI: 10.1101/gad.1418306
This work was funded by a Genomes to Life grant from the U.S. Department of Energy to E.N. and B.T.N. E.N. is a Howard Hughes Medical Institute Investigator. BioCAT is a National Institutes of Health-supported Research Center grant (No. RR-08630). Use of the Advanced Photon Source was supported by the U.S. Department of Energy, Office of Science, Office of Basic Energy Sciences, under Contract No. W-31-109-ENG-38.
Based on an APS press release by Mona Mort.