The element selenium is incorporated into proteins through the 21st amino acid selenocysteine (Sec). Such selenoproteins are critically important to all types of life, suggesting that being able to accurately decode the Sec codon and correctly placing this amino acid in proteins is biologically fundamental. However, little is known about biosynthesis of selenoproteins in eukaryotic cells. To better understand this process, a team of researchers used data gathered at two APS sectors to determine the crystal structure of the human translational elongation factor responsible for recognizing and delivering the transfer rnA (trnA) carrying Sec to the ribosome. They produced some surprising findings, which suggest that the mechanism for elongation factor to incorporate selenium into growing protein chains is different from that of elongation factors associated with every other type of amino acid.
Standard amino acids rely on the elongation factors eef1A and ef-Tu. ef-Tu in particular is made of three sections, known as domains 1, 2, and 3. To perform its job, ef-Tu utilizes a molecule called guanosine triphosphate. Previous research has shown that during the process of delivering an amino acid-carrying (or aminoacyl) trnA into a lengthening protein, one of the phosphates on the gTP molecule ef-Tu carries is cleaved off, or hydrolyzed, which turns it into guanosine diphosphate (gdP). This triggers a major conformational change: domain 1 rotates about 90 degrees away from domains 2 and 3, which helps the trnA release ef-Tu and subsequent positioning at the appropriate site on the ribosome.
However, the trnA associated with Sec requires a different elongation factor. In prokaryotic cells, that factor is Selb, and in eukaryotes, it’s eefSec. Both of these elongation factors are made of four domains. Some studies have shown that Selb doesn’t undergo a conformational change after the gTPto-gdP exchange, but little was known about the behavior of eefSec.
To learn more about this process, investigators from the University of Illinois at Chicago, the Rutgers-Robert Wood Johnson Medical School, the University of Texas Southwestern Medical Center, the Illinois Institute of Technology, and Yale University used the BioCAT 18-id-d, SbcCAT 19-id-d, and LSCAT 21-id beamlines at the APS to determine the crystal structure of eefSec while it’s complexed with non-hydrolyzable analogs of gTP, as well as gdP.
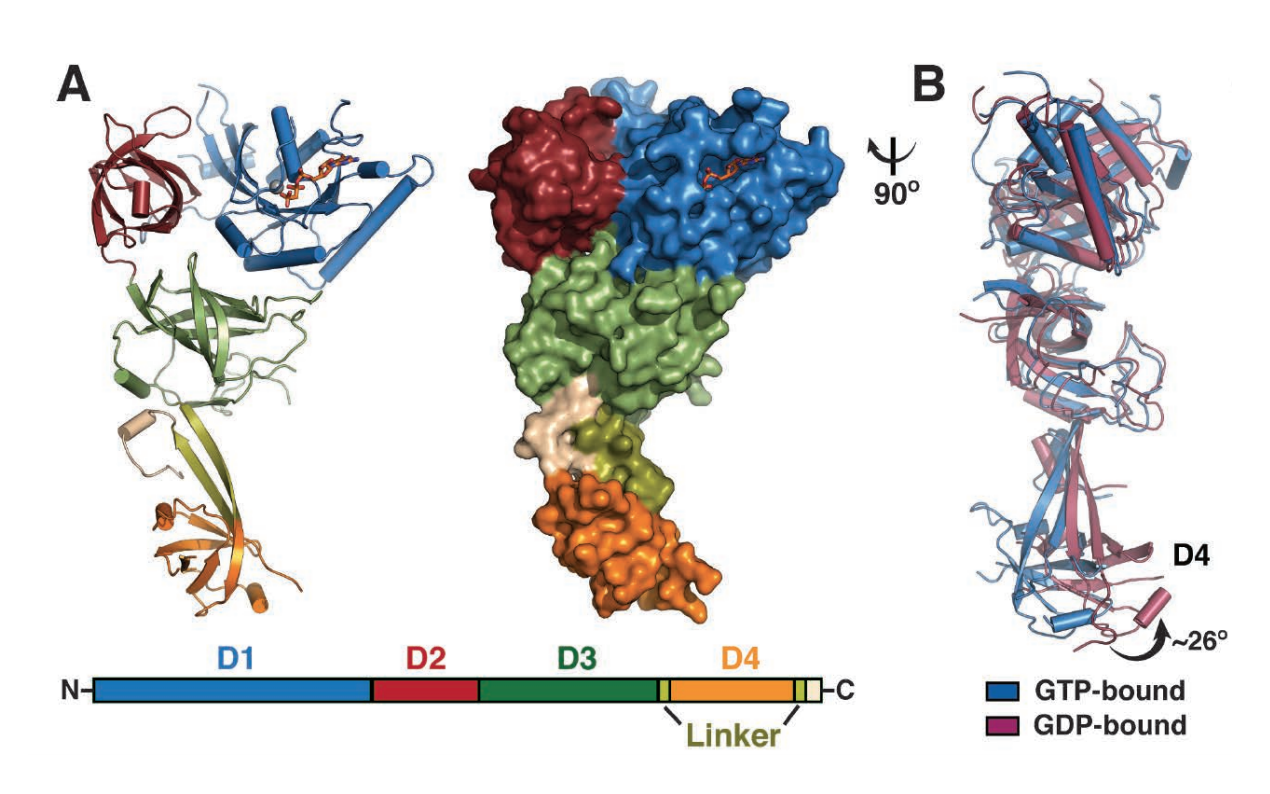
Their findings show that eefSec has a structure shaped like a chalice: domains 1, 2, and 3 represent the cup, a linker region represents the stem, and domain 4 represents the base (fig. 1). Further investigation showed the presence of a hydrogen bond between amino acid residues in domain 3 and the linker that seemed to act like a hinge—a clue that something unusual might take place at this region during hydrolysis.
Sure enough, when the researchers determined the crystal structure of eefSec complexed with gdP, they found that domain 4 swung 26 degrees away from the other three domains. Domains 1 and 2 also underwent small shifts, with domain 1 moving slightly toward the predicted trnA binding face and domain 2 toward the opposite direction.
Although the movement of domains 1 and 2 seem to tighten the Sec-binding pocket, which could help lower the binding affinity of the Sec trnA, the movement of domain 4 is more mysterious. The authors speculate that the swing of domain 4 might help eject the trnA and dissociate eefSec from the ribosome.
More research will be necessary to better define the reasons behind this conformational change, the researchers say. Either way, it differs significantly from the mechanisms that eef1A and ef-Tu use to deliver their aminoacyl-trnAs to growing protein chains. The authors note that another research team recently revealed that some microorganisms use different codons to encode Sec, suggesting that more surprises about this amino acid are in store.
See: Malgorzata Dobosz-Bartoszek, Mark H. Pinkerton, Zbyszek Otwinowski, Srinivas Chakravarthy, Dieter Söll, Paul R. Copeland, and Miljan Simonovic, “Crystal structures of the human elongation factor eefSec suggest a non-canonical mechanism for selenocysteine incorporation,” Nat. Commun. 7, 12941 (2016). doi: 10.1038/ncomms12941
We thank the staff of LSCAT and SbcCAT beamlines at the APS and organizers of the 7th Annual CCP4 USA Crystallography School for their help during x-ray data collection and processing, the staff at the Advanced Protein Characterization Facility (SbcCAT, APS) for access to mosquito crystallization robots, and the staff of BioCAT beamline for help during small-angle x-ray scattering data collection. The initial part of the study was supported by the University of Illinois at Chicago startup fund and a grant from the American Cancer Society, Illinois division (225752 to m.S.). The subsequent studies were supported by grants from the National Institute of General Medical Sciences (gm097042 to m.S., gm070773 to P.r.c. and gm22854 to d.S.). use of LSCAT was supported by the Michigan economic development corporation and the Michigan Technology Tri-corridor (grant 085P1000817). SbcCAT is operated by U. Chicago Argonne, llc, for the U.S. Department of Energy (DOE) Office of Biological and Environmental Research under contract no. de-Ac02-6ch11357. BioCAT is supported by a grant from the National Institute of General Medical Sciences of the National Institutes of Health (P41 gm103622). Use of the Pilatus 3 1M detector was provided by grant 1S10d018090-01 from the National Institute of General Medical Sciences. This research used resources of the Advanced Photon Source, a U.S. DOE Office of Science user facility operated for the DOE Office of Science by ANL under contract no. de-Ac02-06ch11357.
Adapted from an APS press release by Christen Brownlee.