Invariant BECN1 CXXC Motifs Bind Zn2+ and Regulate Structure and Function of the BECN1 Intrinsically Disordered Region
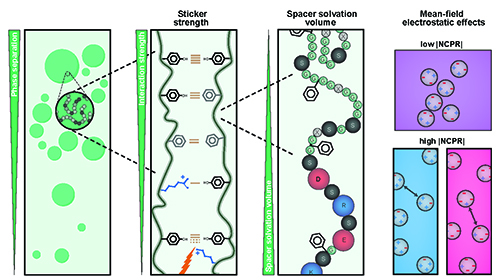
Autophagy is a conserved lysosomal degradation pathway that degrades un-needed cellular components such as misfolded, aggregated, mutated and damaged proteins, organelles, and pathogens. Autophagy dysfunction is implicated in numerous diseases including neurodegenerative disorders, muscular diseases, cardiomyopathy, cancer and infectious diseases. Many proteins involved in autophagy contain intrinsically disordered regions (IDRs) that do not form stable secondary or tertiary structure. The structural flexibility of IDRs is thought to enable diverse and multiple interactions enabling them to regulate cell signaling pathways. Many IDRs have been shown to fold upon binding to ligands. BECN1, a key autophagy protein involved in autophagosome nucleation, contains two invariant CxxC motifs within a large BECN1 intrinsically disordered region (IDR) at the BECN1 N-terminus. The goal of the research was to uncover the functional roles of the invariant CxxC motifs which were hitherto not understood. The authors used inductively coupled plasma mass spectrometry to demonstrate that BECN1 binds Zn2+ in a 1:1 molar ratio, and that mutation of the invariant cysteines prevents co-ordination of Zn2+, demonstrating that the CxxC motifs are responsible for binding Zn2+. Zn2 …